Cytoscape node attributes12/25/2023 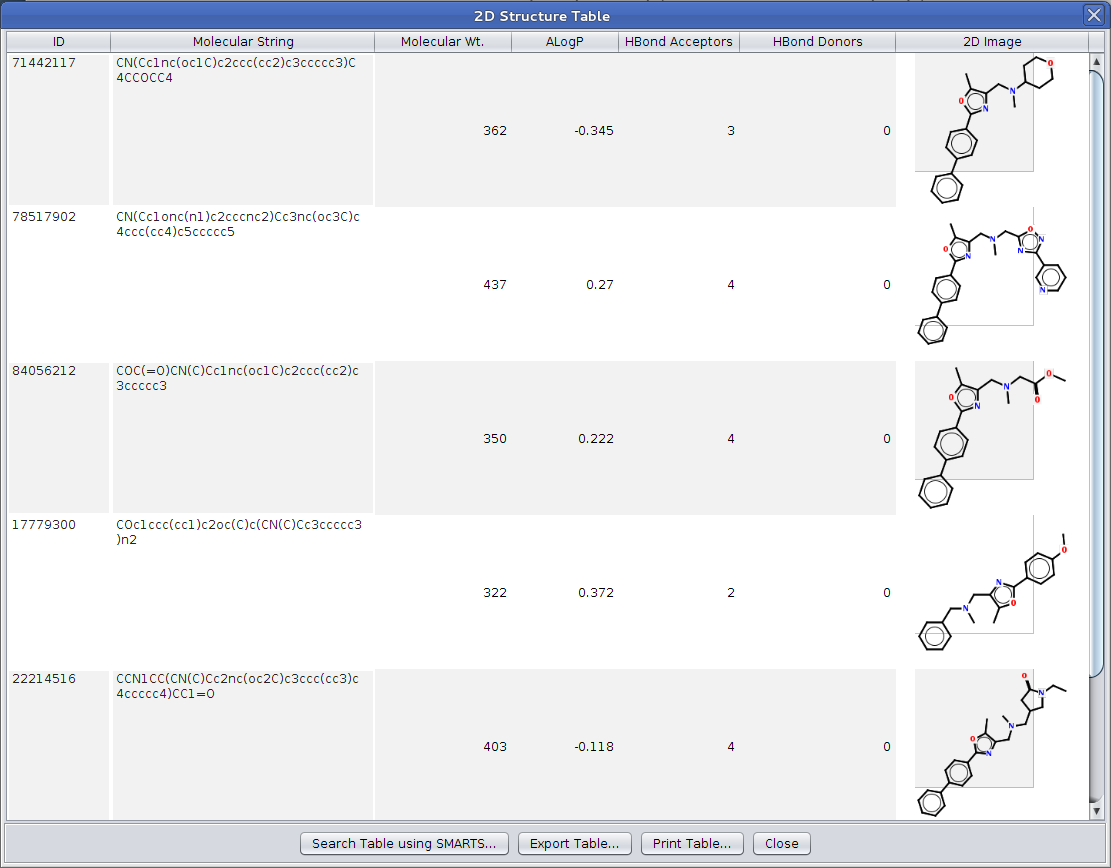
NetMatch computes and displays results graphically on the right side of the window. Users set NetMatch parameters on the left side of the window, then press ‘Go’. The NetMatch main window showing the results of a small 3-node query. Users can express queries in NetMatch by (i) loading from an existing file, (ii) importing from the Cytoscape workspace, (iii) drawing using the NetMatch query drawing tool. NetMatch can be set to interpret labeled/unlabeled, directed/undirected networks ( Fig. The joining process connects the subparts by all paths satisfying the approximate paths present in the query. Such consistency rules allow the pruning of the search space, reducing significantly the computational cost of the matching process.Īpproximate query graphs are handled by first independently processing all maximal specified subparts and then ‘joining’ in all possible ways the results of the subqueries. When a pair of nodes is added to the partial mapping, consistency conditions are checked. The aim of the matching process is the determination of a mapping, which is a bijection, and consists of a set of node pairs covering all the query graph nodes. The searching (matching) process is carried out by using the state space representation (Cordella et al., 2004) where a state is a partial mapping and a transition between two states corresponds to the addition of a new pair of matched graph nodes. NetMatch handles target and query graphs with multi-edges (more than one edge between two nodes), loops (edges starting and ending at the same node) and a list of attributes for each edge and node. Approximate queries are special subgraphs that may contain: (i) nodes and edges labeled with a special wildcard symbol ‘?’, which can match any single value of a user specified node or edge attribute (ii) approximate paths, which are paths of length ≤ n or ≥ n, where n is a positive integer, that can connect two nodes. NetMatch supports subgraph matching queries against a target network, previously loaded into the Cytoscape workspace. NetMatch has been implemented as a plugin for Cytoscape (Shannon et al., 2003), an open source software platform for network visualization and analysis that is extensible through a straightforward plug-in architecture, allowing rapid development of additional features. NetMatch provides an efficient graph matching algorithm (Cordella et al., 2004) with extensions to handle multiple labels per node, multiple edges between pairs of nodes, and approximate queries. The query results are subgraphs of the original graph connected in the same way as the query graph. For instance, the tYNA (Yip et al., 2006) Cytoscape plugin can submit a network to a remote server for detection of four common motifs and the MAVisto (Schreiber and Schwobber-meyer, 2005) and FANMOD (Wernicke and Rasche, 2006) software find over-represented network motifs of a user defined size.Ī query to NetMatch is a graph, some of whose elements are constants and some are wildcards (which can match an unspecified number of elements). 1Ī small number of tools currently exist for querying networks for motifs, but none of these allow user-defined queries implemented as easy to use software. In epidemiology, acquaintance graphs between people may characterize specific patterns of disease outbreaks and can be used to optimize vaccine delivery. In cell biology, it may be of interest to see whether the connectivity of genes of one functional type is similar to some characteristic shape, like a feed-forward loop. Locating subgraphs matching a specific topology is useful to find higher-order connectivity motifs of networks that may have functional relevance in the modeled biological system. 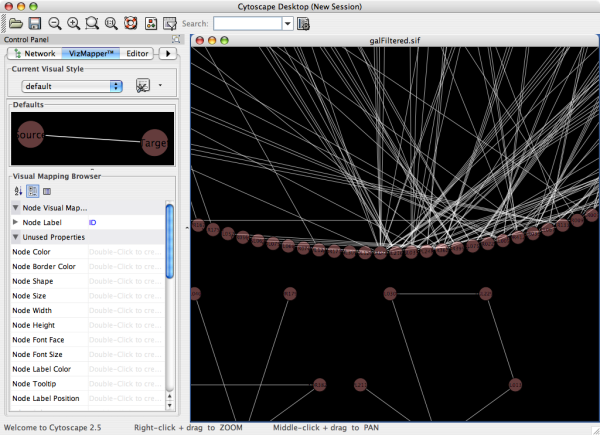
Such networks are naturally modeled as large graphs, which can be analyzed using graph theoretical techniques. Many biological systems arise from complex interactions between components (people, organisms, cells, proteins, DNA, RNA and small molecules) (Barabási and Oltvai, 2004).
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
